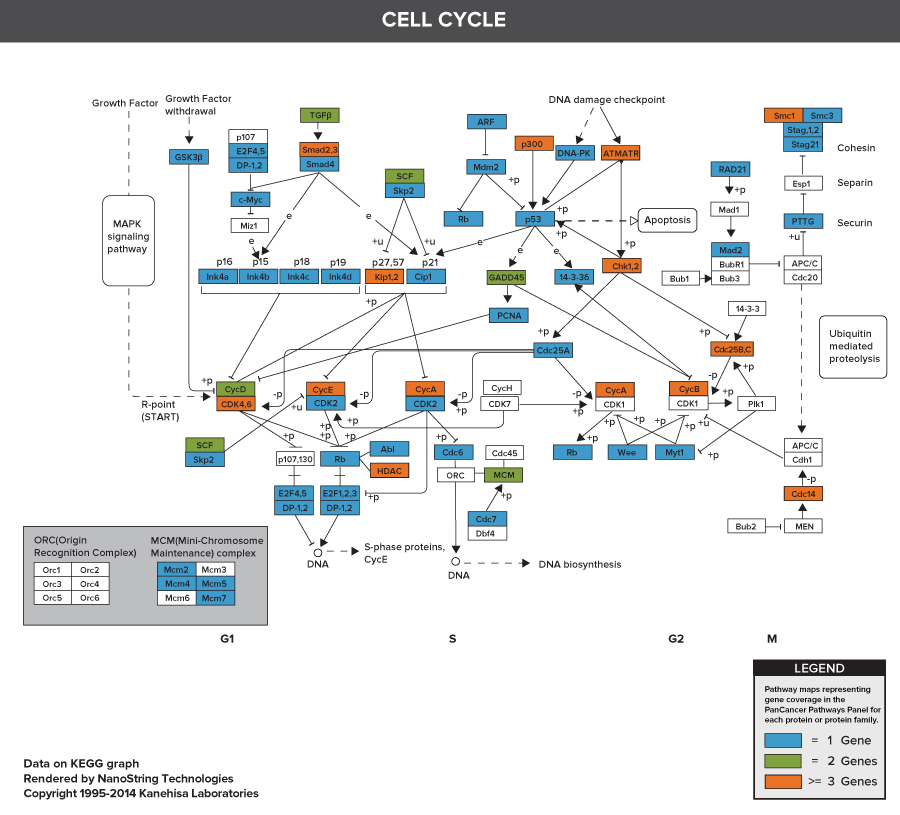
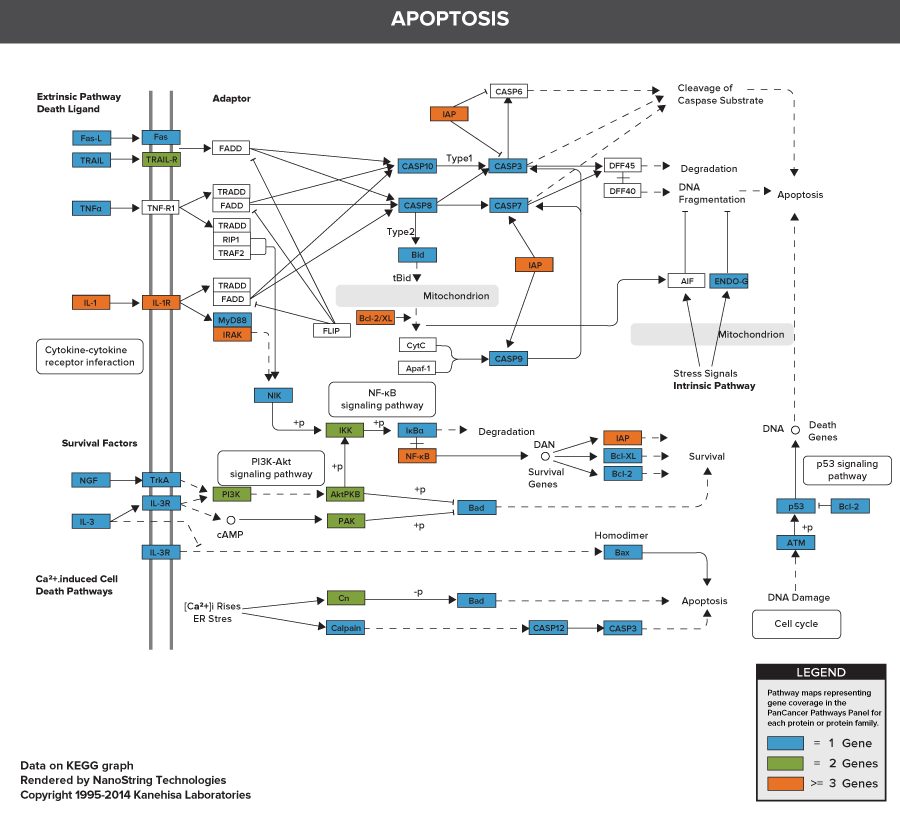
nCounter PanCancer Pathways Panel
Drawing on 30+ years of cancer research, QPCR, Next Generation Sequencing, Gene Expression, Western blotting the cancer community has accomplished a lot to narrow tens of thousands of genes to the ones that matter for the study of cancer.
Today, we are taking the next step in that journey with the introduction of our PanCancer Pathways Panel, a novel set of 700 essential genes representing all major cancer pathways including key driver genes.
Using a biology-guided, data-driven approach and applying basic co-expression principles we scored and ranked each gene based on its biological relevance to cancer and its role as an essential representation in one or more of the 13 canonical pathways.
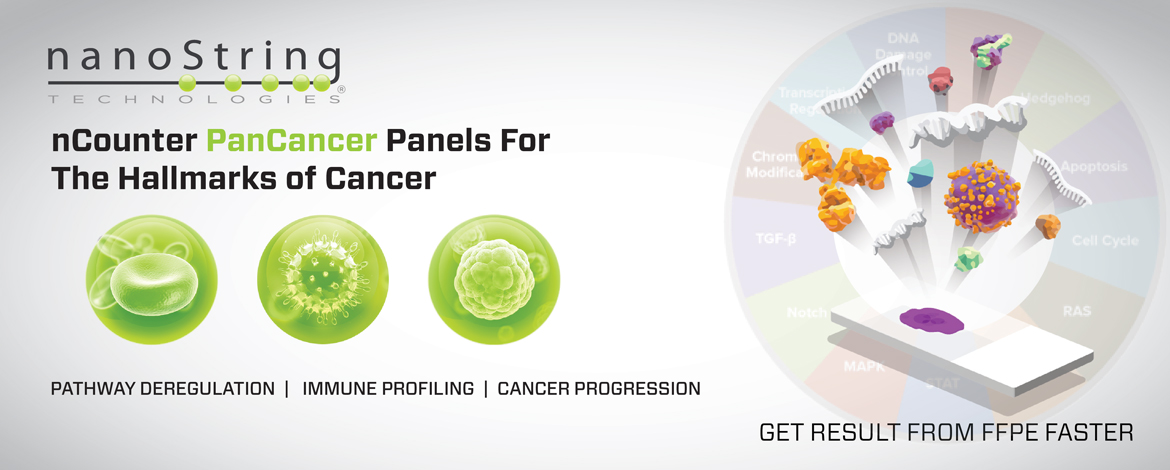
nCounter PanCancer Panels
Our suite of nCounter PanCancer Panels includes content driven by key areas of cancer biology as described in the Hallmarks of Cancer. Key panels include:
|
||
| PanCancer Immune Profiling Panel Explore the immune response to cancer. |
||
| PanCancer Pathways Panel Investigate cancer progression and metastasis. |
Download the Nanostring PanCancer Panels Product Bulletin here.
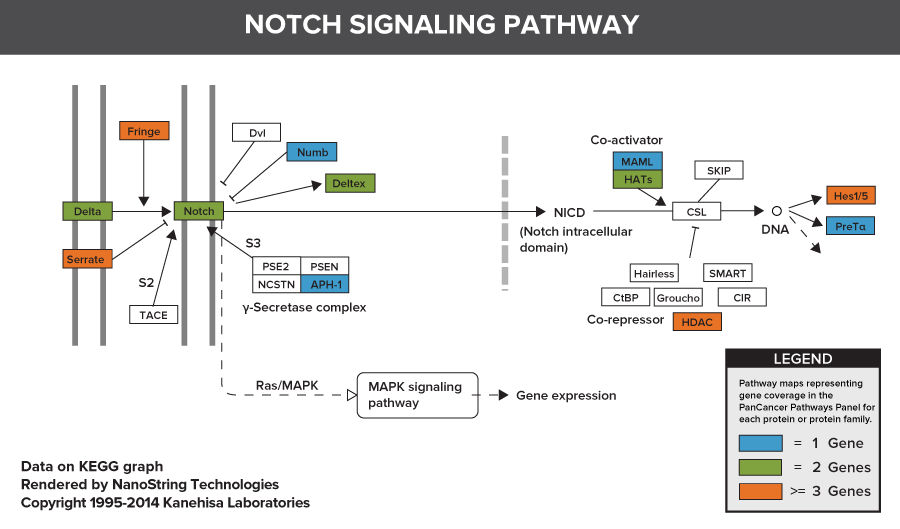
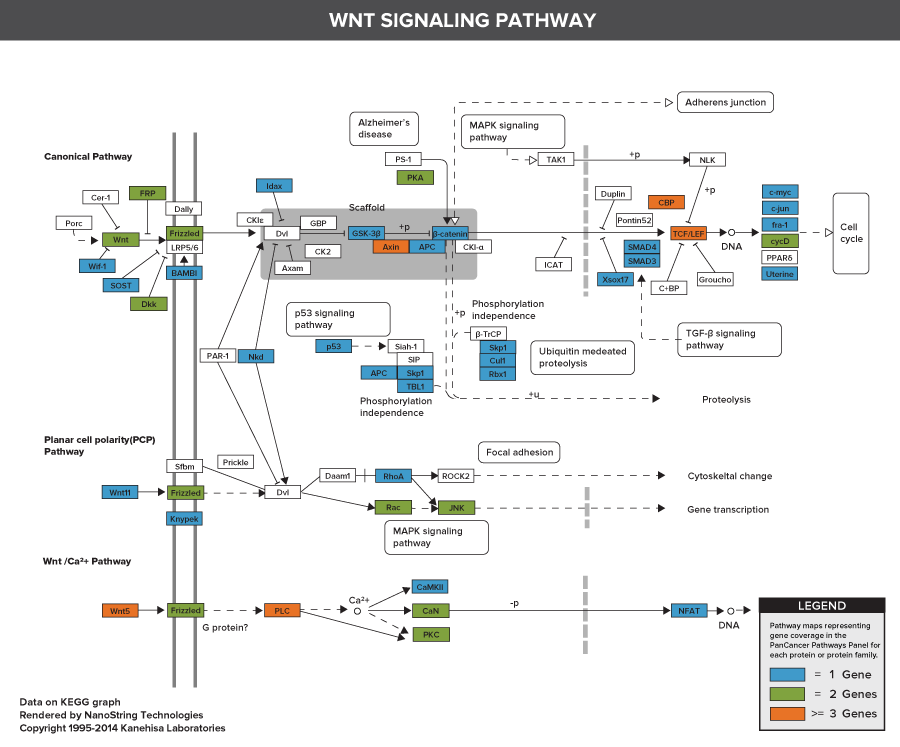
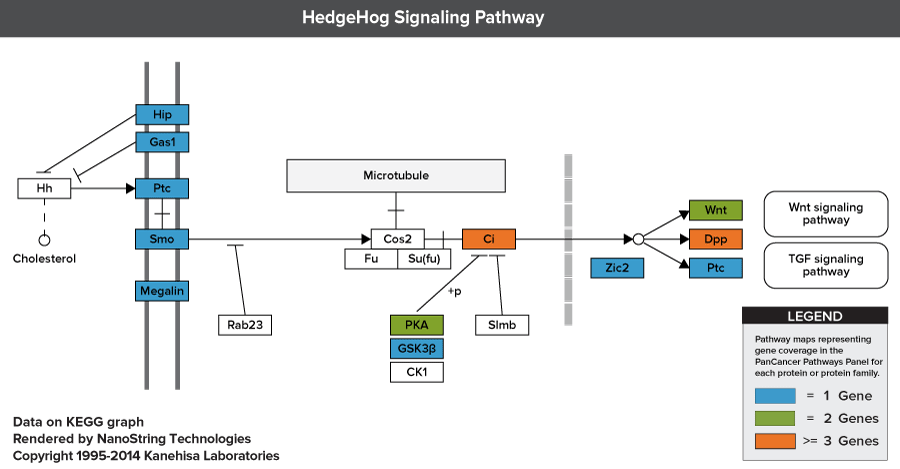
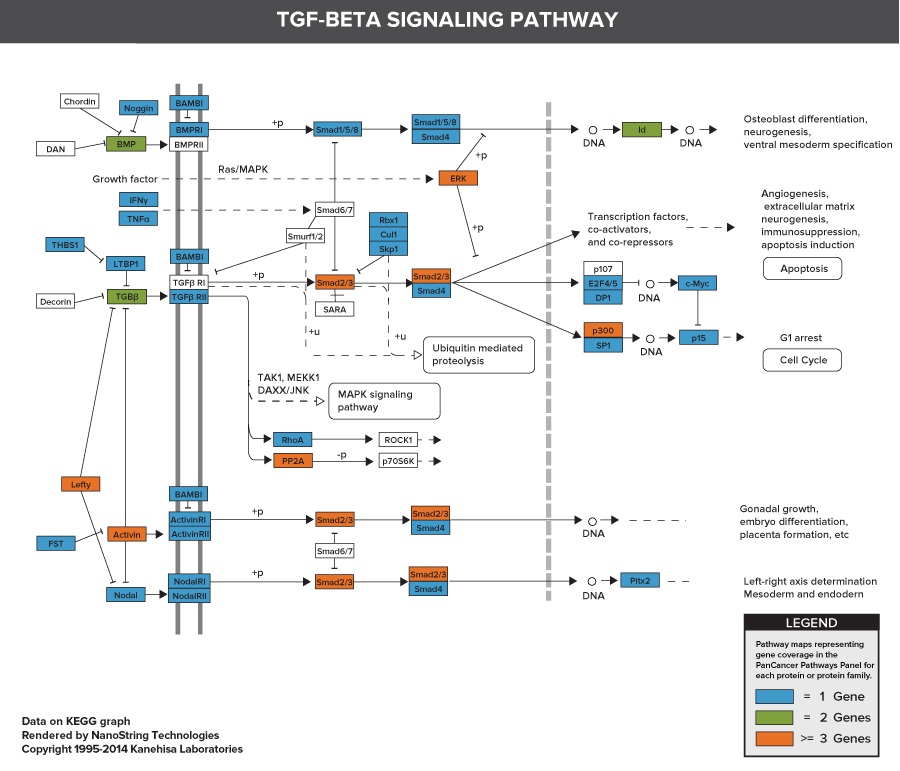
Pathways of the PanCancer Pathways Panel
|
770 Essential Genes Representing 13 Canonical Pathways
|
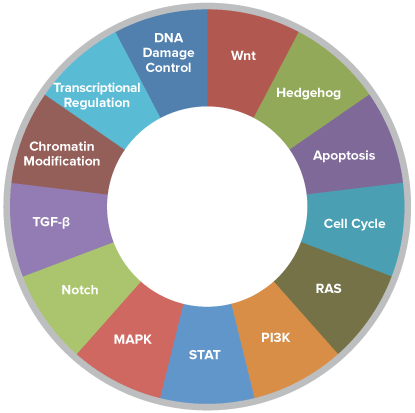 |
Drawing from data generated through TCGA and major thought leaders in the field each gene has been scored and ranked based on it’s biological relevance to cancer and its role as an essential representation of 13 canonical pathways
Complementary Advanced Data Analysis
Don’t waste time waiting for a bioinformatics specialist to review your data. Get results in a simple CSV file that contains direct counts for each target. Import it into your favorite application or use the included nSolver™ Analysis Software for convenient and advanced data analysis.
nSolver™ Analysis Software enables easy storage, custom QC, and custom normalization of nCounter data. Generate highly-customized exports, basic statistical outputs, and publication-quality figures quickly and easily with the included tools. Specialized Advanced Analysis Modules for PanCancer panels, including custom and panels provides deeper data insights based on robust statistics. Examine experimental trends, identify pathway-specific responses, and profile immune cell populations in sharable HTML reports.
Click the banner below to view the example HTML report generated by nSolver:
PanCancer Pathway Description



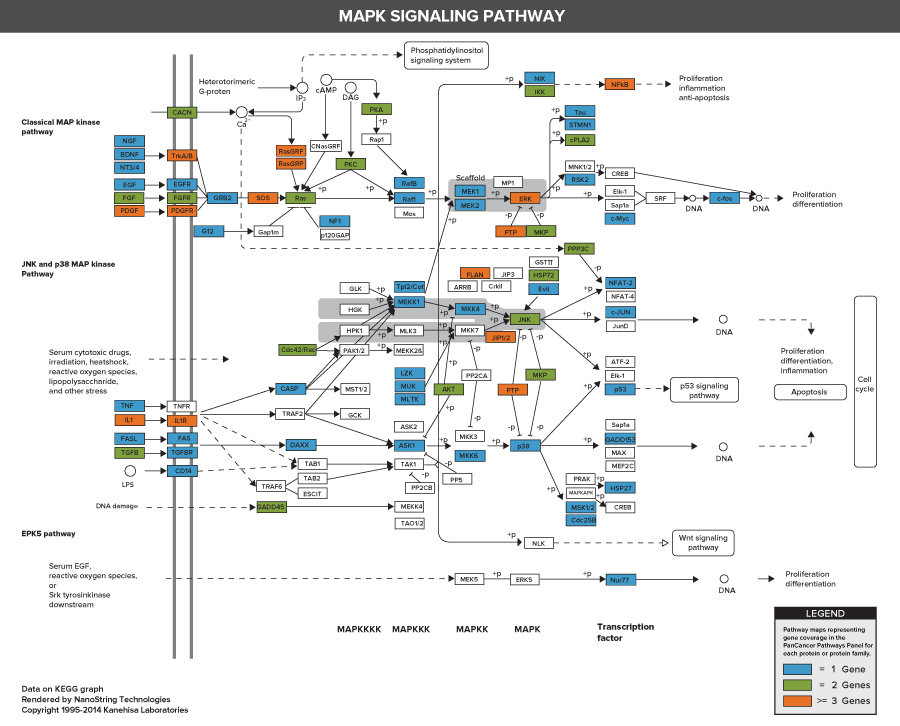
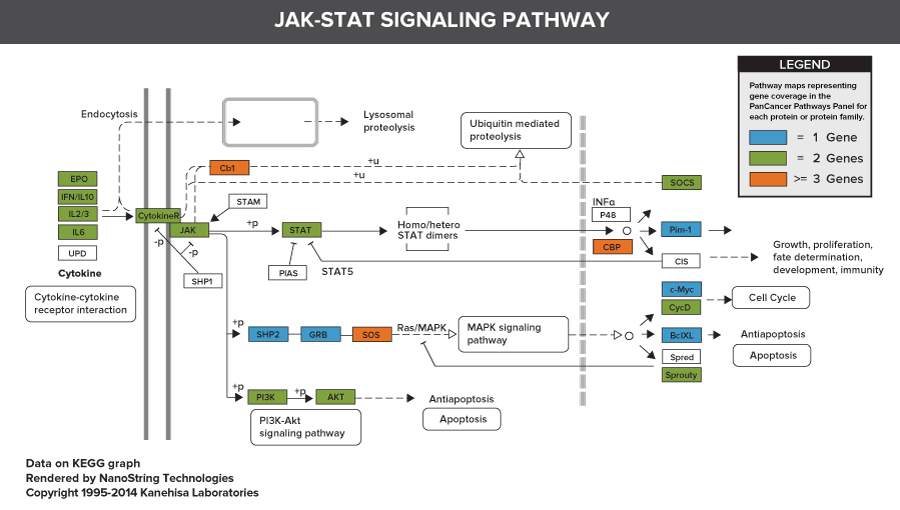
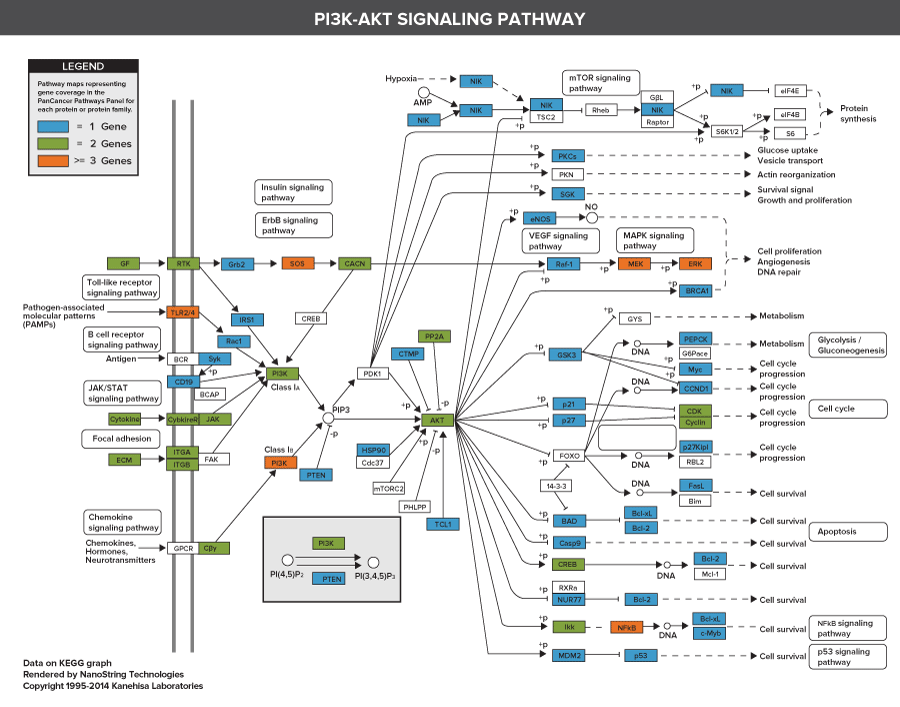
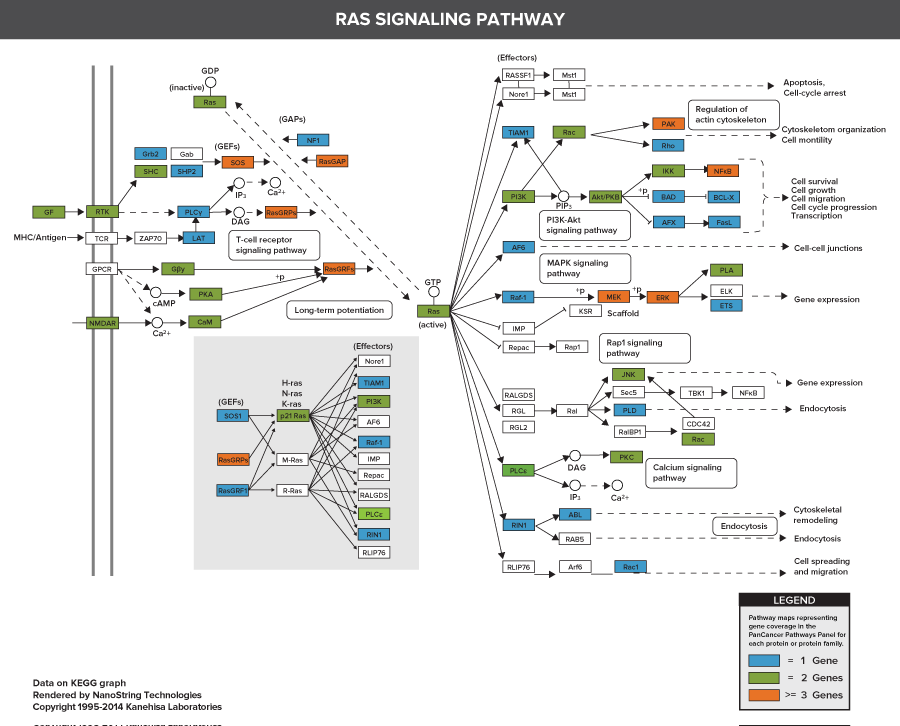
 606 Pathway Genes: Notch, Wnt, Hedgehog, TGFB, MAPK, STAT, P13K, RAS, Chromatin Modification, Transcriptional Regulation, DNA Damage Control, Cell Cycle, Apoptosis
606 Pathway Genes: Notch, Wnt, Hedgehog, TGFB, MAPK, STAT, P13K, RAS, Chromatin Modification, Transcriptional Regulation, DNA Damage Control, Cell Cycle, Apoptosis